A comprehensive review of approaches for genetic improvement in foxtail millet (Setaria italica L.)
Shivika Pareek, Reginah Pheirim*, C.P. Chetariya, Alka Soharu
Department of Genetics and Plant Breeding, School of Agriculture, Lovely Professional University, Jalandhar, Punjab-144411, India
* Corresponding author: Reginah Pheirim (pheirimreginah4@gmail.com)
Abstract: Foxtail millet (Setaria italica (L.) P. Beauv.) is a historically significant and resilient cereal crop known for its adaptability to diverse environmental conditions, nutritional benefits and economic potential. Despite its importance, foxtail millet remains underutilized compared to major cereals. Recent advances in breeding techniques and molecular marker technologies have substantially enhanced efforts toward its genetic improvement. Conventional breeding approaches, including selection, hybridization and mutation breeding, have contributed to trait improvement, while modern strategies such as marker-assisted selection (MAS), genome-wide association studies (GWAS), and quantitative trait loci (QTL) mapping have accelerated the identification of genes associated with desirable agronomic traits. The development of high-resolution genetic maps and the use of molecular markers, including simple sequence repeats (SSRs) and single-nucleotide polymorphisms (SNPs), have further facilitated genomic studies and breeding programmes. In addition, association mapping has emerged as an effective approach for identifying genomic regions linked to important agronomic traits, thereby supporting precision breeding. With the integration of genomics, transcriptomics and genome-editing technologies, foxtail millet improvement is progressing towards enhanced yield, stress tolerance and nutritional quality. This review summarizes recent advances in foxtail millet genetics, breeding strategies and molecular marker technologies, highlighting their significance for sustainable agriculture and global food and nutritional security.
Keywords: Foxtail millet, breeding approaches, molecular marker technologies, MAS, GWAS, genome editing
Introduction
Setaria italica (L.) P. Beauv., commonly known as foxtail millet, is among the earliest domesticated millet species and has a long history of cultivation in India since ancient times (Jia et al, 2013). Known locally as ‘Kangni’ in Hindi, foxtail millet is a hardy crop that is well adapted to a wide range of climatic conditions. It can thrive in both low-fertility and well-drained fertile soils and is commonly cultivated in semi-arid and rainfed regions (Fukunaga and Kawase, 2024). Owing to its ability to grow under low-input conditions, the crop is often regarded as a ‘life-saving crop’ for farmers in marginal environments (Zhang et al, 2022).
Among the minor millets, foxtail millet ranks as the second most important species and is recognized for its high nutritional value. The grain typically contains about 65% carbohydrates, 11% protein and 6% fat, and is rich in minerals such as iron and copper, as well as dietary fibre (Yousaf et al, 2021). It serves as a staple food in some regions of Southern India. Due to its immense nutritional value, millet is also considered one of the best weaning foods. In today's modern world, where growing consumer demand for healthier food, foxtail can serve as one of the best choices for the diet/weight-conscious, diabetic and people with heart disease, and holds great potential due to its unique phenolic content and roughage (Tripathi et al, 2021). In addition to human consumption, foxtail millet is used in the production of alcoholic beverages and also serves as fodder and feed for livestock in both temperate and tropical regions (Tomar et al, 2023).
Despite its resilience, adaptability, and nutritional advantages, foxtail millet remains far less widely cultivated than major cereal crops. Historically, the crop has received limited research attention, leading to its underutilization and its perception as a ‘forgotten crop,’ often labelled as a ‘poor man’s food’ (Singh et al, 2023). In contrast, major cereals such as rice, maize and wheat, which together contribute over 60% of the global plant-based dietary energy supply, are relatively poor sources of essential micronutrients. This heavy reliance on micronutrient-poor staple crops has significantly contributed to the problem of ‘hidden hunger’ (Vetriventhan et al, 2019).
With the rapidly increasing global population and the growing challenges of climate change, there is an urgent need to diversify agricultural systems to ensure sustainable food and nutritional security. In this context, foxtail millet has gained renewed attention due to its climate resilience, nutritional richness and potential role in sustainable agriculture. Therefore, this review provides a comprehensive overview of foxtail millet, with particular emphasis on its genetic resources, breeding strategies and molecular approaches that have been explored for its genetic improvement.
Morphological description of foxtail millet
Foxtail millet is a robust and upright annual grass that typically grows to a height of 0.6–1.2m (Brink, 2006). The plants are mostly slender with stem tillers but may occasionally branch, typically producing 1 to 25 culms, with an average of 3–4. It possesses a well-developed root system, with slender, wiry adventitious roots originating from the lower nodes (Rahayu et al, 1996). Leaves are simple and alternate with a glabrous or slightly hairy sheath measuring 10 to 25cm in length. The ligule is short and fimbriate, while the linear leaf blade varies from 15 to 50cm in length and 0.5 to 4cm in width, featuring an acuminate apex and a slightly rough, prominent midrib. It has a short vegetative growth period (Dekker, 2003), ranging from 80 to 120 days, though some cultivars mature within 60 days (Swamy, 2023).
Floral biology
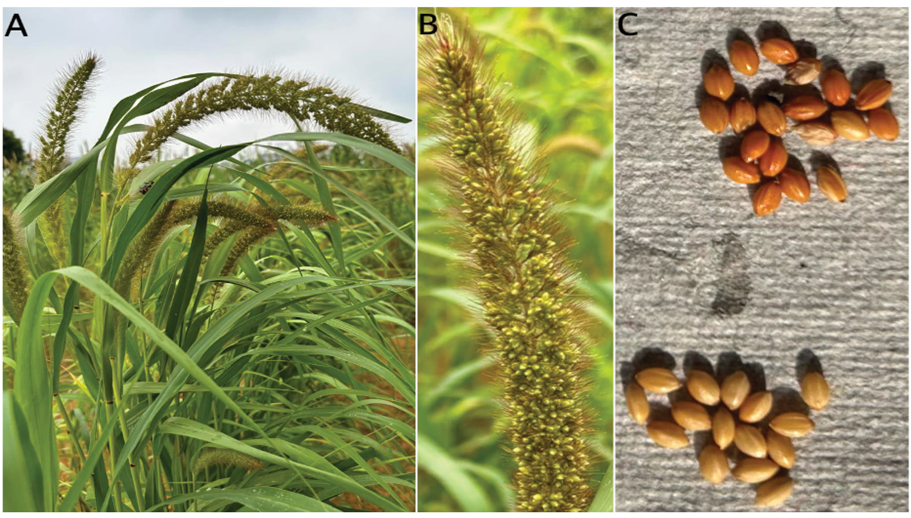
The millet’s inflorescence, measuring 6–30cm, consists of a central stalk surrounded by short, bristling, prickly lateral branches (Figure 1B). The terminal spike usually ranges from 8 to 32cm in length with a drooping appearance (Figure 1A), is thick with cylindrical, lobed structure and held up by a very tiny, slender pedicel (Sundararaj and Thulasidas, 1976). It consists of pairs of tiny blooms on each spikelet, which are enclosed by two glumes. The top bloom is fertile and bisexual, while the lower flower is sterile (Nirmalakumari and Vetriventhan, 2010). Each floret consists of three stamens with anthers that are white or yellow in colour (Jayaraman et al, 1997). The pistil of the flower consist of a smooth, round ovary with two long styles and feathery stigmas. The fruit is a caryopsis (grain), approximately 2mm in length and oval in shape. The colour varies from pale yellow to orange, red, brown, or black, with the lemma and palea tightly enclosing the grain with a 1,000-seed weight of around 2g (Figure 1C).

Anthesis and pollination
Foxtail millet is a self-pollinating species with a low average natural outcrossing rate of approximately 4%, occasionally spontaneous hybrids forming between wild and cultivated varieties (Till-Bottraud et al, 1992). Flowering commences when roughly three-quarters of the head protrudes from the sheath and proceeds to flower in a vertically aligned manner (Sundararaj and Thulasidas, 1976). The flowering process for a head takes between 8 to 16 days. Anthesis commonly occurs early in the morning and around midnight, although this can vary widely based on environmental conditions (Siles et al, 2001).
Global status of germplasm resources
S. italica is predominantly grown in Asia, parts of Europe, and Africa, playing a vital role in the diets of people in China, India, Korea, Japan and Nepal (Dwivedi et al, 2012). Countries such as China, India, France and Japan have the most extensive collections of foxtail millet germplasm (Vetriventhan et al, 2016). The Crop Trust, along with other global organizations, is actively engaged in the ex situ and in situ conservation of foxtail millet genetic resources (Bramel et al, 2022). The Chinese National Gene Bank (CNGB) preserves a vast collection of 26,670 germplasm accessions (Wang et al, 2012). The International Crops Research Institute for the Semi-Arid Tropics (ICRISAT) houses germplasm sourced from 26 different countries, while the Plant Genetic Resources Conservation Unit (PGRCU) in the USA and the National Institute of Agro-biological Sciences (NIAS) in Japan also maintain diverse germplasm collections (Upadhyaya et al, 2009; 2011). The formation of representative germplasm, which includes core and mini core collections (Upadhyaya et al, 2009; 2011), serves as valuable genetic resources for research at the genomic level.
Cytogenetics studies in foxtail millet
Foxtail millet is a C4 crop with a chromosome number of 2n = 2x = 18 (AA), belonging to Panicoid, characterized by a comparatively small genome size of 515Mb approx (Lata et al, 2013). It belongs to the Panicoideae subfamily and the tribe Paniceae. The wild ancestor of cultivated foxtail millet (S. italica) is the green foxtail millet, S. viridis (2n = 2x = 18, AA). Wild species S. faberii and S. verticillata consist of an AABB genome, thought to have arisen from natural crossing between S. viridis and S. adhaerans. A species from Mexico, S. grisebachii, is a diploid species detected to have a CC genome. The sole species that is autotetraploid in the Setaria genus with an AAAA genome was identified in S. queenslandica. Other polyploid species identified in Setaria are S. pumila and S. pallidefusca (Benabdelmouna et al, 2001a; 2001b; Benabdelmouna and Darmency, 2003).
Enhancing crop resilience, production, and adaptation in adverse environmental conditions is largely dependent on genetic diversity. Establishing breeding goals and comprehending the genetic diversity of foxtail millet are important initial steps in developing superior cultivars that can meet the growing global demand (Ramesh et al, 2023). A wide range of genetic variation in the population of foxtail millet has been reported based on traits such as the number of tillers, bristle length, panicle orientation, panicle compactness, anther colour, and seed size (Wang et al, 2012; Moharil et al, 2019). ICRISAT analyzed approximately 1,535 S. italica samples from 26 different countries to look at differences in height of plant, time of blooming, structure of inflorescence, and the shape of seed amongst the collection. The possibility of gene transfer between Setaria species appears to be significant, especially between cultivated and wild species, since the outcrossing rate was shown to range significantly (0.3–4%). The variation in the crop has been explored through several studies, including pedigree, morphological and biochemical assessment within foxtail millet (Murugan and Nirmalakumari, 2006; Nirmalakumari and Vetriventhan, 2010). The biochemical analyses done in foxtail millet were based on seed protein analysis, isozymes (Jusuf and Pernes, 1985), and analysis of molecular markers or DNA markers (Schontz and Rether, 1999; Van et al, 2008; Fukunaga et al, 2002; Jia et al, 2009; Radha et al, 2014). Foxtail millet's genetic diversity is protected and encouraged, which might strengthen our ability to assist international efforts to develop a stronger and fairer food system and to encourage agricultural innovation (Govindaraj et al, 2020).
Breeding approaches in foxtail millet
Improving grain yield, nutritional composition, drought tolerance and resistance to pests and diseases while adapting to evolving demands can be achieved through the strategic use of diverse germplasm in breeding programmes. Below is an overview of breeding approaches employed in foxtail millet improvement (Figure 2).
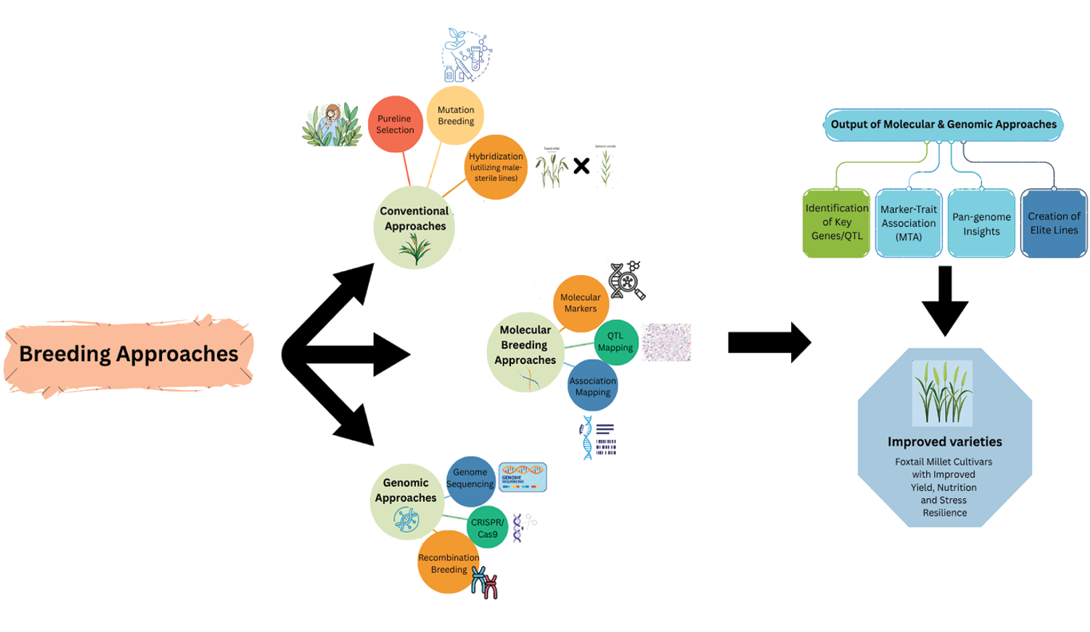
Conventional approaches
In the early stages of foxtail millet breeding, pure line selection was the predominant method for enhancing grain yield. Breeding approaches for foxtail millet encompass selection and hybridization utilizing male-sterile lines (Swamy, 2023). Genetically male sterile lines governed by dominant gene ‘Ch A’ (Hu et al, 1986) and photoperiod-sensitive male sterility (Wensheng et al, 1991) have been developed for heterosis breeding in foxtail millet to facilitate the development of hybrid varieties (Liu et al, 2014). Hybridizing foxtail millet (Setaria italica) with its wild counterpart S. viridis was reported to incorporate triazine resistance into the cultivated species (Darmency and Pernes, 1985). S. viridis acts as an important genetic reservoir for improving foxtail millet, offering a straightforward and efficient breeding strategy (Naciri et al, 1992). The main challenge in conventional breeding is inbreeding depression, which results in the loss of heterosis or hybrid vigour. Another issue is linkage drag, where undesirable genes are transferred and integrated into the genome.
Mutation breeding approaches
Mutation breeding has significantly contributed to crop enhancement on a global scale. Its primary goal is to amplify the frequency and range of mutations while enhancing the occurrence of viable mutations for targeted genetic modifications. Induced mutagenesis acts as a viable alternative breeding strategy to enhance variability and address specific shortcomings in existing cultivars. Various mutant varieties were listed in the MVD (Mutant Variety Database), among which a few were foxtail millet (S. italica L.) resulting from direct or indirect mutation breeding with chemical and physical mutagens (IAEA, 2022). Some examples of mutant varieties released were Lugu 7, developed by irradiation of seeds with gamma rays with improved attributes of 10–15cm shorter stem and resistance to lodging, Jingu 47, officially approved in 2009, developed by treatment in aerospace, with main improved attributes of high yield and good quality. Also, mutagenesis techniques have been effectively combined with modern molecular biology tools, such as molecular marker analysis and high-throughput mutation screening, thereby increasing their efficiency and impact on crop improvement (Shu, 2009). Some examples in foxtail millet are SiDWARF2, a dwarf mutant gene (Xue et al, 2016), SiYGL1, a yellow-green leaf mutant (Li et al, 2016), and siago1b, a gene showing pleiotropic developmental defects (Liu et al, 2016).
Molecular breeding approaches
Conventional breeding often requires a long duration to develop and commercialize the cultivars, and several traits are easily influenced by environmental factors or exhibit low heritability. Advances in molecular biology led to the development of molecular marker technologies, which enable the study of plants based on polymorphic DNA sequences. Molecular markers are easily identified DNA fragments widespread in the genome with no environmental effect, and non-specific to tissue and stages of plants. It provides the most appropriate tool for the genetic diversity assessment, allowing the selection of suitable parental lines for further breeding programmes, efficient handling of plant genetic resources, and identifying varieties (de Vienne, 2003). The molecular approaches in foxtail millet are discussed broadly in the subsequent sections.
Molecular marker technologies
Molecular marker technologies have significantly advanced genetic studies and breeding programmes in foxtail millet, enabling precise identification of genetic variations. These markers are essential for assessing the genetic diversity, trait inheritance and crop improvement using marker-assisted selection (MAS). Among the molecular markers, restriction fragment length polymorphisms (RFLP) were the first molecular markers used in foxtail millet to analyze genetic differentiation across geographical diversity based on heterologous ribosomal DNA probes, providing foundational insights into domestication and diversity of foxtail millet and its wild ancestor S. viridis (Schontz and Rether, 1999; Fukunaga et al, 2002). RFLP map in the crop was constructed from an inter-varietal cross, Longgu 25 × Pagoda Flower green and reported nine linkage groups by Wang et al (1998). Doust et al (2004) expanded the work by using 257 RFLP markers from rice and foxtail millet to identify quantitative trait loci (QTL) in the aspect of branching and inflorescence architecture. Using F2 plants derived from B100 (S. italica) and A10 (S. viridis) as a mapping population, the first functional dissection of morphological characters in foxtail millet using molecular mapping was developed. PCR-based molecular markers such as amplified fragment length polymorphism (AFLP), random amplified polymorphic DNA (RAPD), ISSR markers, and simple sequence repeats (SSRs) are widely used for genetic diversity studies, germplasm characterization, and linkage map construction (Gupta and Varshney, 2004; Jain et al, 2009; Kumari et al, 2011; Ardie et al, 2017). Among the molecular markers, SSR markers have been utilized to a great extent due to their reproducibility, high polymorphism and ease of use. This marker has been most extensively used in foxtail millet facilitating the study of diversity (Chander et al, 2017; Ramesh et al, 2023; Reddy et al, 2025) for linkage map construction (Jia et al, 2009; Sato et al, 2013; Fang et al, 2016) and QTL mapping (Wang et al, 2017; Gao et al, 2025). The development of SSR markers based on the genomic resources of the draft genome in foxtail millet enables germplasm characterization, linkage mapping, phylogenetics and comparative mapping in foxtail millet (Pandey et al, 2013). Transposon display (TD marker) has been employed to examine the genetic structure of foxtail millet and wild green foxtail (S. italica subsp. viridis (L.) P. Beauv.) revealing geographical structuring (Hirano et al, 2011). The advent of single nucleotide polymorphism (SNP) has emerged as a powerful, high-resolution tool in molecular breeding and genomic research. The resequencing-based marker systems offers to detect SNPs and InDels and rapidly transitioned from low throughput to high throughput-based techniques and offers the construction of high-density linkage maps (Wang et al, 2017; Guo et al, 2023), genome wide association study (Jia et al, 2013; Jaiswal et al, 2019; 2024) and fine mapping of QTLs (Du et al, 2021; Li et al, 2022) in foxtail millet.
Methods for identification of QTLs in foxtail millet
Numerous chromosomal regions and QTLs have been identified that regulate a wide range of phenotypic traits in foxtail millet. According to Yu and Buckler (2006), the majority of the desirable features in plant breeding, such as height, quality, resistance to diseases, drought and salinity, are inherently quantitative in nature and are controlled by several genes. Such polygenic traits are regulated by specific genomic regions referred to as quantitative trait loci (QTLs). Phenotypic variations in complex traits arise from the combined effects of environmental factors, multiple QTL influences, QTL-QTL interactions, or QTL–environment interactions (Maloof, 2003). The primary goal of QTL identification is to locate genomic regions associated with complex traits and to identify tightly linked and neutral inherited molecular markers that can be used in breeding programmes (Agarwal et al, 2008). QTL identification involves the sequential arrangement of markers and determining genetic distances, assigning linkage groups based on recombination values. In order to address this complexity, high-resolution and accurate tools are required not only to decipher trait architecture but also to aid in the development of MAS tools for breeding programmes. QTL identification in plants is predominantly carried out using two approaches: (1) QTL mapping and (2) association mapping.
(1) QTL mapping in foxtail millet: Identified and genomic regions
The concept of QTL mapping was first introduced by Karl Sax in 1923. QTL mapping involves the analysis of an appropriate mapping population. Several QTL accounting for agronomic and yield attribute traits were reported in foxtail millet (Doust et al, 2004; Wang et al, 2013; Fang et al, 2016). QTL mapping was employed in the crop for stb1 gene, for ‘spikelet-tipped bristles’ (Sato et al, 2013), variations in blooming time under various environmental circumstances (Mauro-Herrera et al, 2013), and germination and drought tolerance in early planting (Qie et al, 2014). The integration of next-generation sequencing enabled the development of high-density SNP based linkage maps. Map-based cloning was used to fine map and clone SiDWARF2 (a foxtail millet dwarf mutant) derived from Yugu1 (Xue et al, 2016), a recessive nuclear gene SiYGL1 encoding a magnesium-chelatase D subunit, responsible for yellow–green leaf mutant (Li et al, 2016), and Argonaute 1 (AGO1) mutant (siago1b) induced by ethyl methane sulfonate exhibiting pleiotropic developmental defects in foxtail millet (Liu et al, 2016). Liu et al (2024) identified QTLs associated with downy mildew resistance based on the specific locus amplified fragment sequencing with high-density linkage map and the phenotype data in four environments. They also concluded through collinearity analysis between genomes of pearl millet and foxtail millet that the genes were taxon-specific (Liu et al, 2024). Ma et al (2025) identified four important hull colour QTL and provided a basis for characterizing hull colour indices and contributing to the advancement of QTL mapping for grain colour. Liu et al (2022) identified 221 QTLs for 17 morpho-agronomic and yield-related traits, while Yoshitsu et al (2017) identified two QTLs (qDTH2 and qDTH7) regulating days to heading (DTH) using QTL-seq analysis. Ni et al (2017) performed resequencing of 184 recombinant inbred lines (RILs) of foxtail millet for QTL mapping of nine agronomic traits and identified a single gene controlling five traits, two QTLs for plant height and three QTLs for heading date. This provides an efficient way for constructing a high-resolution genome assembly and identifying genes. Han et al (2024) developed a genetic map for plant height based on re-sequencing, identifying 19 unconditional and 13 conditional QTLs. QTLs identified using molecular markers in foxtail millet are shown in Table 1.
Table 1. A summary of QTL identification in foxtail millet. Abbreviations: QTL, quantitative trait loci; RFLP, restriction fragment length polymorphism; SSR, simple sequence repeat; GWAS, genome-wide association study; SNP, single nucleotide polymorphism; GBS, genotyping-by-sequencing; RAD-seq, restriction site-associated DNA sequencing; MTA, marker–trait association; RIL, recombinant inbred line; BSR-seq, bulked segregant RNA sequencing; SLAF-seq, specific length amplified fragment sequencing; InDel, insertion–deletion; ICIM, inclusive composite interval mapping; MCIM, mixed composite interval mapping.
|
Germplasm |
Mapping approach |
Population type |
Marker used |
No. of markers |
Genomic region(s) identified |
Putative gene identified |
Trait(s) studied |
References |
|
Cross between Setaria italica (B100) × S. viridis (A10) |
QTL mapping (Composite Interval Mapping, comparative genomics) |
F2:3 population (~120 families) |
RFLP markers |
119 markers (subset from 257 loci map) |
Multiple QTL across chromosomes I–IX; major regions on chromosomes V, VI, IX |
tb1 (minor effect), auxin & gibberellin pathway genes, monoculm1, dwarf-related genes |
Tillering, axillary branching |
|
|
Diverse germplasm panel |
Association mapping |
Diverse germplasm panel, 184 foxtail millet accessions |
Genomic SSRs |
50 |
Strong associations on chromosome 5 |
SSR b129 (Ubiquitin carboxyl-terminal hydrolase) and SSR p75 (Phospholipid acyltransferase) |
Yield-related agronomic attributes |
|
|
Global Collection |
GWAS Association Mapping) |
181 accessions, including 155 from the core collection |
GBS |
17,714 |
Pigmentation and days to 50% flowering at chromosomes 4 and 6, respectively |
- |
Flowering time and plant pigmentation |
|
|
Segregating population |
Linkage mapping |
F2 population (Yugu1 × Longgu7, 167 individuals) |
SSRs |
10,598 |
QTLs distributed across all nine chromosomes |
- |
Agronomic and yield attributes |
|
|
RILs |
High-density linkage mapping and QTL analysis |
184 RILs (Zhanggu × A2) |
SNPs |
483,414 SNPs; 3,437 recombination bins |
QTLs on all nine chromosomes |
sd1 (gibberellin synthesis gene) |
Agronomic traits |
|
|
Segregating population |
RAD-seq based QTL mapping |
F2 population from cross between Hongmiaozhangu × Changnong35 (124 individuals) |
SNPs |
9,968 |
11 QTLs on chromosome 1,2,5,7,8,9 |
- |
Agronomic traits |
|
|
Yuikogane × Shinanotsubuhime (F2 population, 382 plants) |
QTL-seq combined with Bulked Segregant Analysis (BSA) and Composite Interval Mapping (CIM) |
F2 population |
SNPs, Indel markers, CAPS markers |
~45,370 SNPs identified between parents; 24 Indel & CAPS markers used for validation |
Two major QTLs: qDTH2 (Chr 2: 38.2–39.6Mb) and qDTH7 (Chr 7: 29.2–31.0Mb) |
Candidate genes include Seita.2G286100 (OsPRR95 homolog), Seita.2G291300 (DLF1 homolog) and Seita.7G246700 (Roc4 homolog) |
Days to Heading (DTH) / Flowering time |
|
|
Cultivars |
GWAS Association Mapping |
142 diverse genotypes (core collection) |
SNPs (GBS-ddRAD) |
12,460 |
High-confidence MTAs on chromosomes 3, 6, 7, 9 |
27 candidate genes for 1,000-grain weight, grain yield and flag leaf width |
Major agronomic traits |
|
|
Yugu1 × Longgu7 |
QTL mapping (linkage mapping using resequencing data) |
RIL population |
Bin markers + SSR markers |
2297 bin markers + 74 SSR markers |
221 QTL across genome; 22 QTL clusters; stable QTL qLMS6.1 (Chr 6) |
Seita.6G250500 |
Morpho-agronomic and yield-related traits (plant height, tiller number, panicle traits, yield traits) |
|
|
Aininghuang × Jingu21 |
QTL mapping (unconditional & conditional QTL analysis + transcriptome integration) |
RIL population |
Bin markers (resequencing-based) |
4360 bin markers |
19 unconditional QTL + 13 conditional QTL; 4 stable QTL across environments |
8 candidate genes (identified via RNA-seq & WGCNA; specific names not mentioned in abstract) |
Plant height (PH) |
|
|
Foxtail millet G1 × JG21 |
QTL mapping using high-density linkage map and BSR-seq validation |
Recombinant Inbred Lines (F6:7, 158 lines) |
SLAF-seq SNP bin markers |
1031 bin markers |
Major region on Chr8 (0.78 Mb interval) including qDM8_1, qDM8_2, qDM8_4 |
Seita8G.199800, Seita8G.195900, Seita8G.198300, Seita8G.199300 (NBS-LRR genes) |
Downy mildew resistance |
|
|
Foxtail millet Jingu28 × Ai88 |
QTL mapping using genetic linkage map |
F2 population (300 individuals) |
SSR + InDel markers |
215 (213 SSR + 2 InDel) |
Major QTLs qHD9-1 (Chr9) and qPH5-1 (Chr5); 46 QTLs forming 13 clusters |
Seita.9G020100 (CCT motif gene), Seita.5G404900 (GA20-oxidase) |
12 agronomic traits including heading date and plant height |
|
|
Foxtail millet Changsheng07 × Donggu218 |
QTL mapping using genetic linkage map |
F2 population with F2:3 families |
SSR + InDel markers |
196 (159 SSR + 37 InDel) |
Major QTLs qMPL3.1, qMPL5, qMPW2, qSD5, qTGW5.1, qTGW5.2, qGL5; 22 QTLs forming 4 clusters |
- |
Panicle-related traits (panicle length, width, grain length, TGW) |
|
|
Yugu18 × Hongjiugu19 |
QTL mapping (ICIM, MCIM, multi-method phenotyping) |
RIL (F6, 250 lines) |
SNPs, InDels, bin markers |
~20,748 SNPs + 1,759 InDels (1420 bins) |
Major QTL: qHC1.1, qHC1.2 (Chr 1); qHC9.1, qHC9.3 (Chr 9) |
Not specifically identified (QTL regions validated; overlap with previous loci) |
Hull colour (grain colour traits using 4 methods) |
(2) Advances in association mapping or association analysis in foxtail millet
Association mapping or association analysis uses linkage disequilibrium (LD) to investigate the connection between genotypic constitution and phenotypic expression in natural populations (Borba et al, 2010). It is typically conducted using two approaches – candidate gene-based and genome-wide association mapping. The major advantages of this approach are its use of natural populations, the elimination of bi-parental population requirements, and its ability to achieve high-resolution mapping due to recombination events accumulated over multiple generations (Agrama et al, 2007). GWAS was performed using the core collection of foxtail millet as an association mapping panel, which involves genome-wide screening of nucleotide sequence variation. Gupta et al 2014, evaluated 50 SST markers on 184 foxtail millet accessions across nine chromosomes and identified eight significant markers linked to agronomic traits and two markers with significant association for ubiquitin carboxyl-terminal hydrolase and phospholipid acyltransferase. GWAS by Upadhyaya et al (2015) identified several SNP loci associated, as well as a major genomic region of plant pigmentation and flowering time. The genomic regions of plant pigmentation were identified between 7.2 and 7.3Mbp on chromosome 4, which was also reported by Jia et al (2013) for the pigmentation-related traits such as colour of bristle, leaf sheath and pulvinus. Significant associations were identified on chromosome 6, specifically around 34.0 to 35.5Mbp.
Advances in genomic research for foxtail millet improvement
Among minor millets, foxtail millet possesses the smallest genome (423–510Mb) and was the first millet crop to have its entire genome sequenced. Its compact diploid genome, rapid growth cycle and self-pollination make it a model for C4 species (Vetriventhan et al, 2020). Zhang et al (2012) made a major breakthrough, producing a draft genome (~423Mb) anchored to nine chromosomes. This provides marker discovery, gene annotation and comparative genomics with other cereals such as rice, pearl millet and sorghum. The reference genome allowed the development of a genome-wide scale of microsatellite markers across the nine chromosomes of foxtail millet. A diverse foxtail millet collection was genotyped through genotyping-by-sequencing (GBS) by ICRISAT in collaboration with Cornell University, identifying genome-wide SNPs and assessing population structure and diversity (Upadhyaya et al, 2015). Several gene families have been identified and characterised in foxtail millet, including NAC (NAM, ATAF, and CUC) (Puranic et al, 2013), MYB (myeloblastosis transcription factor) (Muthamilarasan et al, 2014b), WRKY (WRKY DNA-binding protein) (Muthamilarasan et al, 2015), AP2/ERF (APETALA2/Ethylene-Responsive Factor) (Lata et al, 2014), and C2H2 (Cys2-His2 zinc finger protein) (Muthamilarasan et al, 2014a), which play important roles in stress response, growth, and development. In addition, regulatory components such as DCL (Dicer-like), AGO (Argonaute), and RDR (RNA-dependent RNA polymerase) (Yadav et al, 2015) are involved in gene silencing pathways. Although these gene families are conserved across plant species, their characterisation in foxtail millet provides insights into mechanisms of stress tolerance and can aid genetic improvement.
The Beijing Genomics Institute, China and The Joint Genome Institute, USA, have recently sequenced the complete genome of two foxtail millet accessions. The comparative genome mapping through sequence alignment of foxtail millet demonstrated a strong syntenic relationship with both rice and sorghum, even though they have diverged over half a century years ago. With this genome-wide sequence resources and availability in public repositories, it is now possible to develop in silico molecular markers and large-scale validation to utilize them in various applications for genetic improvement in foxtail millet. Such a genome-wide scale may prove beneficial for other underutilized or orphan crop species with limited or no genomic information available.
Application of genome editing technologies in foxtail millet
Genome editing technologies have developed as a powerful tool because of their precision and simplicity in modifying targeted genomic regions by using engineered nucleases such as zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TAL-ENs), or RNA-guided engineered nucleases such as the CRISPR/Cas system. The first genome editing in foxtail millet was successfully demonstrated by Lin et al (2018) through mutating the SiPDS gene, which was achieved through protoplast transfection. Cheng et al (2021) targeted gene SiMTL by using CRISPR/Cas9 to generate a haploid inducer line. Although there are relevant reports of genome editing systems in foxtail millet, a highly efficient genome-editing system is limited. Liang et al (2022) first successfully applied base editing in foxtail millet by using CRISPR/Cas9 targeting the SiALS gene, and successfully created a homozygous mutant plant that is herbicide-tolerant.
Future directions and challenges
Foxtail millet is a significant crop with its climatic resilience and superior nutritional profile. With a potential role in food and nutritional security, it is necessary to harness its nutritional potential to meet daily needs for a healthy lifestyle. Addressing the challenges for breeding foxtail millets is necessary to accomplish this goal. An in-depth understanding of the floral biology of foxtail millet is necessary for developing standardized hybridization techniques to create variability, develop improved varieties and enhance tolerance to adverse climatic conditions, which shall serve the breeding efforts of the crop. Exploration of underutilized germplasm, including wild relatives, shall also enhance biotic and abiotic stresses and nutritional quality. The use of advanced genomic tools (e.g. marker-assisted selection and genome-wide association studies) has improved foxtail millet breeding. Collaboration among genebanks, researchers and policymakers is crucial for sustainable utilization of foxtail millet genetic resources (Vetriventhan et al, 2020).
Conclusion
Foxtail millet holds immense potential as a climate-resilient, nutrient-rich crop that can address global food security challenges. Advances in breeding methodologies, including molecular marker technologies and genomic selection, have enabled precise identification and introgression of desirable traits. QTL mapping and association studies have identified key genetic loci governing yield, stress tolerance, and disease resistance, laying the foundation for marker-assisted breeding. The availability of high-throughput sequencing and genome-editing tools such as CRISPR/Cas9 has further enhanced the scope of foxtail millet improvement. However, challenges such as limited genetic diversity in cultivated varieties, underutilization of wild germplasm, and the need for better characterization of genomic resources persist. Further research should prioritize integrating advanced genomic tools, promoting international germplasm exchange and developing climate-resilient cultivars. By leveraging genetic innovations and sustainable breeding approaches, foxtail millet can play an important role in uplifting global agricultural productivity and ensuring food security amid climate change.
Author contributions
Shivika Pareek: Compilation of the literature, preparation of the draft and preparation of figures/tables; Reginah Pheirim: Conceptualization and article revision; Chetariya Chana Pitha: Revision of article; Alka Soharu: Revision of article.
Conflict of interest statement
The authors have no relevant financial or non-financial competing interests to disclose.
References
Agarwal, M., Shrivastava, N., Padh, H. (2008). Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Reports, 27(4), 617-631. https://doi.org/10.1007/s00299-008-0507-z
Agrama, H. A., Eizenga, G. C., Yan, W. (2007). Association mapping of yield and its components in rice cultivars. Molecular Breeding, 19(4), 341-356. https://doi.org/10.1007/s11032-006-9066-6
Ardie, S., Khumaida, N., Fauziah, N., Yudiansyah, Y. (2017). Biodiversity assessment of foxtail millet (Setaria italica L.) genotypes based on RAPD marker. J. Trop. Crop Sci. 4:21–25. https://doi.org/10.29244/jtcs.4.1.21-25
Benabdelmouna, A., Darmency, H. (2003). Copia-like retrotransposons in the genus Setaria: Sequence heterogeneity, species distribution and chromosomal organization. Plant Systematics and Evolution, 237(3), 127-136. https://doi.org/10.1007/s00606-002-0250-9
Benabdelmouna, A., Abirached-Darmency, M., Darmency, H. (2001a). Phylogenetic and genomic relationships in Setaria italica and its close relatives based on the molecular diversity and chromosomal organization of 5S and 18S-5.8 S-25S rDNA genes. Theoretical and Applied Genetics, 103(5), 668-677. https://doi.org/10.1007/s001220100596
Benabdelmouna, A., Shi, Y., Abirached-Darmency, M., & Darmency, H. (2001b). Genomic in situ hybridization (GISH) discriminates between the A and the B genomes in diploid and tetraploid Setaria species. Genome, 44(4), 685-690. https://doi.org/10.1139/g01-032
Borba, T. C. D. O., Brondani, R. P. V., Breseghello, F., Coelho, A. S. G., Mendonça, J. A., Rangel, P. H. N., Brondani, C. (2010). Association mapping for yield and grain quality traits in rice (Oryza sativa L.). Genetics and Molecular Biology, 33, 515-524. https://doi.org/10.1590/S1415-47572010005000065
Bramel, P., Giovannini, P., Eshan Dulloo, M. (2022). Global strategy for the conservation and use of genetic resources of selected millets. Global Crop Diversity Trust, Bonn, 85.
Brink, M. (2006). Setaria italica (L.) P. Beauv. Cereals and Pulses, Plant Resources of Tropical Africa.
Chander, S., Bhat, K. V., Kumari, R., Sen, S., Gaikwad, A. B., Gowda, M. V. C., Dikshit, N. (2017). Analysis of spatial distribution of genetic diversity and validation of Indian foxtail millet core collection. Physiology and Molecular Biology of Plants, 23(3), 663-673. https://doi.org/10.1007/s12298-017-0448-5
Cheng, Z., Sun, Y., Yang, S., Zhi, H., Yin, T., Ma, X., Sui, Y. (2021). Establishing in planta haploid inducer line by edited SiMTL in foxtail millet (Setaria italica). Plant Biotechnology Journal, 19(6), 1089. https://doi.org/10.1111/pbi.13584
Darmency, H., Pernes, J. (1985). Use of wild Setaria viridis (L.) Beauv. to improve triazine resistance in cultivated S. italica (L.) by hybridization. Weed Research, 25(3), 175-179. https://doi.org/10.1111/j.1365-3180.1985.tb00633.x
de Vienne, D. (Ed.). (2003). Molecular markers in plant genetics and biotechnology. CRC Press.
Dekker, J. (2003). The foxtail (Setaria) species-group. Weed science, 51(5), 641-656. https://doi.org/10.1614/P2002-IR
Doust, A. N., Devos, K. M., Gadberry, M. D., Gale, M. D., Kellogg, E. A. (2004). Genetic control of branching in foxtail millet. Proceedings of the National Academy of Sciences, 101(24), 9045-9050. https://doi.org/10.1073/pnas.0402892101
Du, X., Wang, Z., Han, K., Lian, S., Li, Y., Zhang, L., Guo, E., Wang, J. (2021). Fine mapping of qPH9, a major quantitative trait locus, responsible for plant height in foxtail millet [Setaria italica (L.) P. Beauv.]. Molecular Breeding, 41(12), 77. https://doi.org/10.1007/s11032-021-01261-w
Dwivedi, S. L., Upadhyaya, H. D., Senthilvel, S., Hash, C. T., Fukunaga, K., Diao, X., Prasad, M. (2012). Millets: genetic and genomic resources. Plant Breed Rev. 2012; 35:247–375.
Fang, X., Dong, K., Wang, X., Liu, T., He, J., Ren, R., Zhang, L., Liu, R., Liu, X., Li, M., Huang, M., Zhang, Z., Yang, T. (2016). A high-density genetic map and QTL for agronomic and yield traits in foxtail millet [Setaria italica (L.) P. Beauv.]. BMC Genomics 17:336. https://doi.org/10.1186/s12864-016-2628-z
Fukunaga, K., Kawase, M. (2024). Crop evolution of foxtail millet. Plants, 13(2), 218. https://doi.org/10.3390/plants13020218
Fukunaga, K., Wang, Z., Kato, K., Kawase, M. (2002). Geographical variation of nuclear genome RFLPs and genetic differentiation in foxtail millet, Setaria italica (L.) P. Beauv. Genetic Resources and Crop Evolution, 49(1), 95-101. https://doi.org/10.1023/A:1013852007770
Gao, L., Zhu, Q., Li, H., Wang, S., Fan, J., Wang, T., Yang, G. (2025). Construction of a genetic linkage map and QTL mapping of the agronomic traits in Foxtail millet (Setaria italica). BMC Genomics, 26(1), 152. https://doi.org/10.1186/s12864-024-11169-2
Govindaraj, M., Rai, K. N., Kanatti, A., Upadhyaya, H. D., Shivade, H., & Rao, A. S. (2020). Exploring the genetic variability and diversity of pearl millet core collection germplasm for grain nutritional traits improvement. Scientific Reports, 10(1), 21177. https://doi.org/10.1038/s41598-020-77818-0
Guo, S., Chai, S., Guo, Y., Shi, X., Han, F., Qu, T., ... & Yang, P. (2023). Mapping of major QTL and candidate gene analysis for hull colour in foxtail millet (Setaria italica (L.) P. Beauv.). BMC Genomics, 24(1), 458. https://doi.org/10.1186/s12864-023-09517-9
Gupta, P. K., Varshney, R. K. (2004). Cereal genomics: an overview. Cereal genomics, 1-18.
Gupta, S., Kumari, K., Muthamilarasan, M., Parida, S. K., Prasad, M. (2014). Population structure and association mapping of yield contributing agronomic traits in foxtail millet. Plant Cell Reports, 33(6), 881-893. https://doi.org/10.1007/s00299-014-1564-0
Han K, Wang Z, Shen L, Du X, Lian S, Li Y, Li Y, Tang C, Li H, Zhang L, Wang J (2024) Mapping of dynamic quantitative trait loci for plant height in a RIL population of foxtail millet (Setaria italica L.). Frontiers in Plant Science, 15, 1418328. https://doi.org/10.3389/fpls.2024.1418328
Hirano, R., Naito, K., Fukunaga, K., Watanabe, K. N., Ohsawa, R., Kawase, M. (2011). Genetic structure of landraces in foxtail millet (Setaria italica (L.) P. Beauv.) revealed with transposon display and interpretation to crop evolution of foxtail millet. Genome, 54(6), 498-506.
https://doi.org/10.1139/g11-015
Hu, H. K., Ma, S. Y., Shi, Y. H. (1986). The discovery of a dominant male-sterile gene in millet (Setaria italica). Acta Agronomica Sinica 12, 73-78
IAEA (2022). Mutant Variety Database. https://mvd.iaea.org/
Jain, S. M., Brar, D. S., Ahloowalia, B. S. (Eds.). (2009). Molecular techniques in crop improvement. Dordrecht: Springer Netherlands.
Jaiswal, V., Bandyopadhyay, T., Singh, R. K., Gahlaut, V., Muthamilarasan, M., Prasad, M. (2024). Multi-environment GWAS identifies genomic regions underlying grain nutrient traits in foxtail millet (Setaria italica). Plant Cell Reports, 43(1), 6. https://doi.org/10.1007/s00299-023-03127-1
Jaiswal, V., Gupta, S., Gahlaut, V., Muthamilarasan, M., Bandyopadhyay, T., Ramchiary, N., Prasad, M. (2019). Genome-wide association study of major agronomic traits in foxtail millet (Setaria italica L.) using ddRAD sequencing. Scientific Reports, 9(1), 5020. https://doi.org/10.1038/s41598-019-41602-6
Jayaraman N, Suresh S, Nirmala A, Ganeshan NM (1997) Genetic enhancement and breeding strategies in small millets. In: National Seminar on Small Millets, 23-24 April, 1997, Coimbatore, India, pp 19-21 (Extended summaries)
Jia, G., Huang, X., Zhi, H., Zhao, Y., Zhao, Q., Li, W., Han, B. (2013). A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nature Genetics, 45(8), 957-961. https://doi.org/10.1038/ng.2673
Jia, X., Zhang, Z., Liu, Y., Zhang, C., Shi, Y., Song, Y., Li, Y. (2009). Development and genetic mapping of SSR markers in foxtail millet [Setaria italica (L.) P. Beauv.]. Theoretical and Applied Genetics, 118(4), 821-829. https://doi.org/10.1007/s00122-008-0942-9.
Jusuf, M., Pernes, J. (1985). Genetic variability of foxtail millet (Setaria italica P. Beauv.) Electrophoretic study of five isoenzyme systems. Theoretical and Applied Genetics, 71(3), 385-391. https://doi.org/10.1007/BF00251177
Kumari, R., Dikshit, N., Sharma, D., Bhat, K. V. (2011). Analysis of molecular genetic diversity in a representative collection of foxtail millet (Setaria italica (L.) P. Beauv.) from different agro-ecological regions of India. Physiology and Molecular Biology of Plants, 17(4), 363–374. https://doi.org/10.1007/s12298-011-0085-3
Lata, C., Gupta, S., Prasad, M. (2013). Foxtail millet: a model crop for genetic and genomic studies in bioenergy grasses. Critical reviews in biotechnology, 33(3), 328-343. https://doi.org/10.3109/07388551.2012.716809
Lata, C., Mishra, A. K., Muthamilarasan, M., Bonthala, V. S., Khan, Y., Prasad, M. (2014). Genome-wide investigation and expression profiling of AP2/ERF transcription factor superfamily in foxtail millet (Setaria italica L.). PLoS One, 9(11), e113092. https://doi.org/10.1371/journal.pone.0113092
Li, W., Tang, S., Zhang, S., Shan, J., Tang, C., Chen, Q., Diao, X. (2016). Gene mapping and functional analysis of the novel leaf color gene SiYGL1 in foxtail millet [Setaria italica (L.) P. Beauv]. Physiologia Plantarum, 157(1), 24-37. https://doi.org/10.1111/ppl.12405
Li, Y., Wu, S., Cao, Y. (1995). Cluster analysis of an international collection of foxtail millet (Setaria italica (L.) P. Beauv.). Euphytica, 83(1), 79-85. https://doi.org/10.1007/BF01677864
Li, F., Niu, J., Yu, X., Kong, Q., Wang, R., Qin, L., Chen, E., Yang, Y., Liu, Z., Lang, L., Zhang, H., Wang, H., Guan, Y. (2022). Isolation and identification of SiCOL5, which is involved in photoperiod response, based on the quantitative trait locus mapping of Setaria italica. Frontiers in Plant Science, 13, 969604. https://doi.org/10.3389/fpls.2022.969604
Li, Y., Wu, S. (1996). Traditional maintenance and multiplication of foxtail millet (Setaria italica (L.) P. Beauv.) landraces in China. Euphytica, 87(1), 33-38. https://doi.org/10.1007/BF00022961
Li, H., Zhu, Q., Wang, T., Zhao, Y., Fan, J., Yang, L., Yang, G. (2025). Genetic linkage map construction and QTL mapping for panicle-related traits in foxtail millet (Setaria Italica (L.) P. Beauv.). Molecular Breeding, 45(9), 72. https://doi.org/10.21203/rs.3.rs-5367279/v1
Liang, Z., Wu, Y., Ma, L., Guo, Y., Ran, Y. (2022). Efficient genome editing in Setaria italica using CRISPR/Cas9 and base editors. Frontiers in Plant Science, 12, 815946. https://doi.org/10.3389/fpls.2021.815946
Lin, C. S., Hsu, C. T., Yang, L. H., Lee, L. Y., Fu, J. Y., Cheng, Q. W., Shih, M. C. (2018). Application of protoplast technology to CRISPR/Cas9 mutagenesis: from single-cell mutation detection to mutant plant regeneration. Plant Biotechnology Journal, 16(7), 1295-1310. https://doi.org/10.1111/pbi.12870
Liu, X., Tang, S., Jia, G., Schnable, J. C., Su, H., Tang, C., Diao, X. (2016). The C-terminal motif of SiAGO1b is required for the regulation of growth, development and stress responses in foxtail millet (Setaria italica (L.) P. Beauv). Journal of Experimental Botany, 67(11), 3237-3249. https://doi.org/10.1093/jxb/erw135
Liu, Z., Zhang, T., Li, C., Bai, G. (2014). Genetic diversity and classification of cytoplasm of Chinese elite foxtail millet [Setaria italica (L.) P. Beauv.] germplasm. Crop Science, 54(2), 659-666. https://doi.org/10.2135/cropsci2012.11.0646
Liu, T., He, J., Dong, K., Wang, X., Zhang, L., Ren, R., Zhang, Z. (2022). Genome-wide identification of quantitative trait loci for morpho-agronomic and yield-related traits in foxtail millet (Setaria italica) across multi-environments. Molecular Genetics and Genomics, 297(3), 873-888. https://doi.org/10.1007/s00438-022-01894-2
Liu, X., Zhang, N., Sun, Y., Fu, Z., Han, Y., Yang, Y., Jia, J., Hou, S., Zhang, B. (2024). QTL mapping of downy mildew resistance in foxtail millet by SLAF-seq and BSR-seq analysis. Theoretical and Applied Genetics, 137(7), 168. https://doi.org/10.1007/s00122-024-04673-2
Ma, Z., Chai, S., Wu, Y., Li, Y., Han, H., Song, H., Gao, J., Feng, B., Yang, P. (2025). QTL identification of hull color for foxtail millet [Setaria italica (L.) P. Beauv.] through four phenotype identification strategies in a RIL population. Seeds, 4(1), 10. https://doi.org/10.3390/seeds4010010
Maloof, J. N. (2003). QTL for plant growth and morphology. Current opinion in plant biology, 6(1), 85-90. https://doi.org/10.1016/S1369526602000080
Mauro-Herrera, M., Wang, X., Barbier, H., Brutnell, T. P., Devos, K. M., Doust, A. N. (2013). Genetic control and comparative genomic analysis of flowering time in Setaria (Poaceae). G3: Genes| Genomes| Genetics, 3(2), 283-295. https://doi.org/10.1534/g3.112.005207.
Moharil, M. P., Ingle, K. P., Jadhav, P. V., Gawai, D. C., Khelurkar, V. C., Suprasanna, P. (2019). Foxtail millet (Setaria italica L.): potential of smaller millet for future breeding. Advances in Plant Breeding Strategies: Cereals: Volume 5, 133-163. https://doi.org/10.1007/978-3-030-23108-8_4
Murugan R, Nirmalakumari A (2006) Genetic divergence in foxtail millet (Setaria italica (L.) Beauv.). Indian J Genet 66(4):339–340
Muthamilarasan, M., Bonthala, V. S., Khandelwal, R., Jaishankar, J., Shweta, S., Nawaz, K., Prasad, M. (2015). Global analysis of WRKY transcription factor superfamily in Setaria identifies potential candidates involved in abiotic stress signaling. Frontiers in Plant Science, 6, 910. https://doi.org/10.3389/fpls.2015.00910
Muthamilarasan, M., Bonthala, V. S., Mishra, A. K., Khandelwal, R., Khan, Y., Roy, R., Prasad, M. (2014a). C2H2 type of zinc finger transcription factors in foxtail millet define response to abiotic stresses. Functional & Integrative Genomics, 14(3), 531-543. https://doi.org/10.1007/s10142-014-0383-2
Muthamilarasan, M., Khandelwal, R., Yadav, C. B., Bonthala, V. S., Khan, Y., Prasad, M. (2014b). Identification and molecular characterization of MYB transcription factor superfamily in C4 model plant foxtail millet (Setaria italica L.). PLoS One, 9(10), e109920. https://doi.org/10.1371/journal.pone.0109920
Naciri, Y., Darmency, H., Belliard, J., Dessaint, F., Pernès, J. (1992). Breeding strategy in foxtail millet, Setaria italica (LP Beauv.), following interspecific hybridization. Euphytica, 60(2), 97-103. https://doi.org/10.1007/BF00029664
Ni, X., Xia, Q., Zhang, H., Quan, Z. (2017). Updated foxtail millet genome assembly and gene mapping of nine key agronomic traits by resequencing a RIL population. GigaScience, 6(2), 1–8. https://doi.org/10.1093/gigascience/giw005
Nirmalakumari, A., Vetriventhan, M. (2010). Characterization of foxtail millet germplasm collections for yield contributing traits. Electronic Journal of Plant Breeding, 1(2), 140-147.
Pandey, G., Misra, G., Kumari, K., Gupta, S., Parida, S. K., Chattopadhyay, D., Prasad, M. (2013). Genome-wide development and use of microsatellite markers for large-scale genotyping applications in foxtail millet (Setaria italica (L.)). DNA Research, 20(2), 197–207. https://doi.org/10.1093/dnares/dst002
Puranik, S., Sahu, P. P., Mandal, S. N., B, V. S., Parida, S. K., Prasad, M. (2013). Comprehensive genome-wide survey, genomic constitution and expression profiling of the NAC transcription factor family in foxtail millet (Setaria italica L.). PloS One, 8(5), e64594. https://doi.org/10.1371/journal.pone.0064594
Qie, L., Jia, G., Zhang, W., Schnable, J., Shang, Z., Li, W., Diao, X. (2014). Mapping of quantitative trait locus (QTLs) that contribute to germination and early seedling drought tolerance in the interspecific cross Setaria italica× Setaria viridis. PloS One, 9(7), e101868. https://doi.org/10.1371/journal.pone.0101868
Radha, B. N., Channakeshava, B. C., Bhanuprakash, K. (2014). Biochemical landmarks for Identification of foxtail millet genotypes. Intl J Appl, 9(1), 9-26.
Rahayu, M., Jansen, P. C. M. (1996). Setaria italica (L.) P. Beauvois cv. group foxtail millet. Plant Resources of South-East Asia, (10), 127-130.
Ramesh, P., Juturu, V. N., Yugandhar, P., Pedersen, S., Hemasundar, A., Yolcu, S., Chandra Sekhar, A. (2023). Molecular genetics and phenotypic assessment of foxtail millet (Setaria italica (L.) P. Beauv.) landraces revealed remarkable variability of morpho-physiological, yield, and yield-related traits. Frontiers in Genetics, 14, 1052575. https://doi.org/10.3389/fgene.2023.1052575
Reddy, M. P., Rai, V. P., Reddy, C. C. M., Kumar, R., Paliwal, R., Sinha, B. (2025). Genetic diversity and population structure analyses using hypervariable microsatellite markers in foxtail millets (Setaria species): Future smart nutri-cereal crop. South African Journal of Botany, 177, 329-337. https://doi.org/10.1016/j.sajb.2024.11.043
Sato, K., Mukainari, Y., Naito, K., Fukunaga, K. (2013). Construction of a foxtail millet linkage map and mapping of spikelet-tipped bristles 1 (stb1) by using transposon display markers and simple sequence repeat markers with genome sequence information. Molecular Breeding, 31(3), 675-684. https://doi.org/10.1007/s11032-012-9825-5
Schontz, D., Rether, B. (1999). Genetic variability in foxtail millet, Setaria italica (L.) P. Beauv.: identification and classification of lines with RAPD markers. Plant Breeding, 118(2), 190-192. https://doi.org/10.1046/j.1439-0523
Shu, Q. Y. (2009). A summary of the international symposium on induced mutations in plants. In: Shu QY (ed) Induced plant mutations in the genomics era. FAO, UN, Rome, pp 15–18.
Siles, M. M., Baltensperger, D. D., Nelson, L. A. (2001). Technique for artificial hybridization of foxtail millet [Setaria italica (L.) Beauv.]. Crop Science, 41(5), 1408-1412. https://doi.org/10.2135/cropsci2001.4151408x
Singh, D., Lawrence, K., Marker, S., Bhattacharjee, I., Lawrence, R., Choudhary, R., Karunakaran, R. (2023). Rainfed assessment of foxtail millet (Setaria italica L. beauv) germplasms through genotyping and principal component analysis. Frontiers in Plant Science, 14, 1017652. https://doi.org/10.3389/fpls.2023.1017652
Sundararaj DP, Thulasidas G (1976) Botany of Field Crops, Macmillan Publisher, India, 509 pp
Swamy K.R.M. (2023). “Origin, domestication, taxonomy, botanical description, genetics and cytogenetics, genetic diversity, breeding of foxtail millet (Setaria italica (L.) Beauv.)”. International Journal of Development Research, 13, (09), 63660-63682.
Till-Bottraud, I., Reboud, X., Brabant, P., Lefranc, M., Rherissi, B., Vedel, F., Darmency, H. (1992). Outcrossing and hybridization in wild and cultivated foxtail millets: consequences for the release of transgenic crops. Theoretical and Applied Genetics, 83(8), 940-946. https://doi.org/10.1007/BF00232954
Tomar, A., Mishra, A. K., Singh, S. P., Khan, H. H., Patel, P., Singh, R. P. (2023). Nutritional benefit of small millets: Food security and sustainability in India. Pharma Innov J, 12, 3028-33.
Tripathi, M. K., Mohapatra, D., Jadam, R. S., Pandey, S., Singh, V., Kumar, V., Kumar, A. (2021). Nutritional composition of millets. Millets and Millet Technology, 101-119. https://doi.org/10.1007/978-981-16-0676-2_5
Upadhyaya, H. D., Pundir, R. P. S., Gowda, C. L. L., Reddy, V. G., Singh, S. (2009). Establishing a core collection of foxtail millet to enhance the utilization of germplasm of an underutilised crop. Plant Genetic Resources, 7(2), 177-184. https://doi.org/10.1017/S1479262108178042
Upadhyaya, H. D., Ravishankar, C. R., Narasimhudu, Y., Sarma, N. D. R. K., Singh, S. K., Varshney, S. K., Gowda, C. L. L. (2011). Identification of trait-specific germplasm and developing a mini core collection for efficient use of foxtail millet genetic resources in crop improvement. Field Crops Research, 124(3), 459-467. https://doi.org/10.1016/j.fcr.2011.08.004
Upadhyaya, H. D., Vetriventhan, M., Deshpande, S. P., Sivasubramani, S., Wallace, J. G., Buckler, E. S., Ramu, P. (2015). Population genetics and structure of a global foxtail millet germplasm collection. The Plant Genome, 8(3), plantgenome2015-07. https://doi.org/10.3835/plantgenome2015.07.0054
Van, K., Onoda, S., Kim, M. Y., Kim, K. D., Lee, S. H. (2008). Allelic variation of the Waxy gene in foxtail millet [Setaria italica (L.) P. Beauv.] by single nucleotide polymorphisms. Molecular Genetics and Genomics, 279(3), 255-266. https://doi.org/10.1007/s00438-007-0310-5
Vetriventhan, M., Upadhyaya, H. D., Dwivedi, S. L., Pattanashetti, S. K., Singh, S. K. (2016). Finger and foxtail millets. In Genetic and genomic resources for grain cereals improvement (pp. 291-319). Academic press. https://doi.org/10.1016/B978-0-12-802000-5.00007-1
Vetriventhan, M., Upadhyaya, H. D. (2019). Variability for productivity and nutritional traits in germplasm of kodo millet, an underutilised nutrient-rich climate smart crop. Crop Science, 59(3), 1095-1106. https://doi.org/10.2135/cropsci2018.07.0450
Vetriventhan, M., Azevedo, V. C., Upadhyaya, H. D., Nirmalakumari, A., Kane-Potaka, J., Anitha, S., Tonapi, V. A. (2020). Genetic and genomic resources, and breeding for accelerating improvement of small millets: current status and future interventions. The Nucleus, 63(3), 217-239. https://doi.org/10.1007/s13237-020-00322-3
Wang, C., Jia, G., Zhi, H., Niu, Z., Chai, Y., Li, W., Diao, X. (2012). Genetic diversity and population structure of Chinese foxtail millet [Setaria italica (L.) Beauv.] landraces. G3: Genes| Genomes| Genetics, 2(7), 769-777. https://doi.org/10.1534/g3.112.002907
Wang, X. Y., Diao, X. M., Wang, J. Z., Wang, C. F., Wang, G. Q., Hao, X. F., Zhao, F. F. (2013). Construction of Millet Genetic Map and QTL Analysis. Journal of Plant Genetic Resources, 14(5), 871-878.
Wang, Z. M., Devos, K. M., Liu, C. J., Wang, R. Q., Gale, M. D. (1998). Construction of RFLP-based maps of foxtail millet, Setaria italica (L.) P. Beauv. Theoretical and Applied Genetics, 96(1), 31-36. https://doi.org/10.1007/s001220050705
Wang, J., Wang, Z., Du, X., Yang, H., Han, F., Han, Y., Guo, E. (2017). A high-density genetic map and QTL analysis of agronomic traits in foxtail millet [Setaria italica (L.) P. Beauv.] using RAD-seq. PloS One, 12(6), e0179717. https://doi.org/10.1371/journal.pone.0179717
Wensheng, C., Yuzhen, K., Gui, D. (1991). A preliminary observation on breeding of photoperiod-sensitive and dominant GMS (Genic Male-Sterile) material of foxtail millet" Guang A1". Acta Agriculturae Boreali-sinica (China), 6, 47-52.
Xue, C., Zhi, H., Fang, X., Liu, X., Tang, S., Chai, Y., Diao, X. (2016). Characterization and fine mapping of SiDWARF2 (D2) in foxtail millet. Crop Science, 56(1), 95-103. https://doi.org/10.2135/cropsci2015.05.0331
Yadav, C. B., Muthamilarasan, M., Pandey, G., Prasad, M. (2015). Identification, characterization and expression profiling of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in foxtail millet. Plant Molecular Biology Reporter, 33(1), 43-55. https://doi.org/10.1007/s11105-014-0736-y
Yoshitsu, Y., Takakusagi, M., Abe, A., Takagi, H., Uemura, A., Yaegashi, H., Terauchi, R., Takahata, Y., Hatakeyama, K., Yoko, S. (2017). QTL-seq analysis identifies two genomic regions determining the heading date of foxtail millet, Setaria italica (L.) P. Beauv. Breeding Science, 67(5), 518–527. https://doi.org/10.1270/jsbbs.17061
Yousaf, L., Hou, D., Liaqat, H., Shen, Q. (2021). Millet: A review of its nutritional and functional changes during processing. Food Research International, 142, 110197. https://doi.org/10.1016/j.foodres.2021.110197
Yu, J., Buckler, E. S. (2006). Genetic association mapping and genome organization of maize. Current Opinion in Biotechnology, 17(2), 155-160. https://doi.org/10.1016/j.copbio.2006.02.003
Zhang, G., Liu, X., Quan, Z., Cheng, S., Xu, X., Pan, S., Wang, J. (2012). Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nature Biotechnology, 30(6), 549-554. https://doi.org/10.1038/nbt.2195
Zhang, W., Wang, B., Liu, B., Chen, Z., Lu, G., Ge, Y., Bai, C. (2022). Trait selection for yield improvement in foxtail millet (Setaria italica Beauv.) under climate change in the North China plain. Agronomy, 12(7), 1500. https://doi.org/10.3390/agronomy12071500